Protein druggability prediction
April 19, 2007The availability of structural data, especially of proteins complexed to small molecule ligands, has enabled numerous analyses that attempt to understand and predict the forces that govern molecular recognition and ligand binding. Successful drug development requires a disease target that plays a vital role in the causation and progression of the disease phenotype and that can be modulated with a drug molecule.
One approach for evaluating protein druggability is to analyze the genome on the basis of sequence homology to known therapeutic targets.
Another approach which rely on on the 3D structure of the protein target. Systematic analyses of protein srufaces in the search for binding pockets that have high potential to bind small drug like compuunds with high affinity.
To identify all possible binding sites on a protien surface based on algorithms broadly classified into
(i) Geometry-based
Tools based on geometry based alogirthm
(ii)Energy based algorithms
Tools based on Energy based algorithms
- GRID
- vdW-FT
- Drugsite

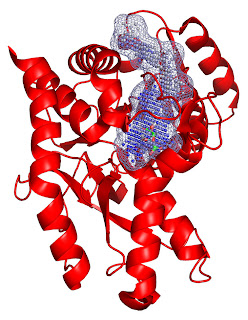 Binding site
Binding site